por Leslie Oros 6 anos atrás
307
old one
por Leslie Oros 6 anos atrás
307

Mais informações
AAUAAA cut the RNA transcript free from the polymerase and releases the pre-mRNA.
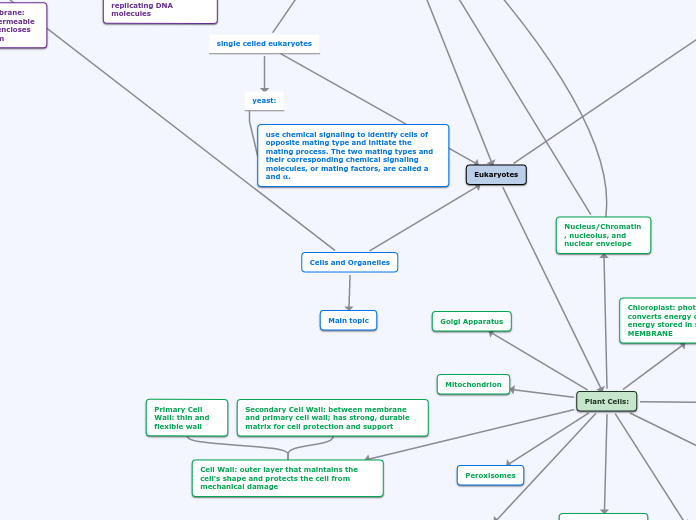
RNA Processing
The ends of the pre-mRNA are modified. The 5' end is synthesized first and receives a 5' cap. The 5' cap is a modified form of a guanine nucleotide.
Polyadenylate polymerase adds 50-250 Adenine nucleotides to the 3' end of the pre-mRNA to make a poly-A tail.
RNA polymerase binds to the promoter (upstream)of a gene, unwinding the DNA to use the strand going from 3' to 5' as the template strand.
Transcription Initiation Complex: when transcription factors and RNA polymerase become bound to the promoter.
Direction of transcription = downstream The other direction = upstream
#1 INITIATION: RNA polymerase initiates RNA synthesis at the starting point on the template strand.
Smooth ER: outer surface, lacks ribosomes, functions in metabolic processes (synthesis of lipids, metabolism of carbohydrates, detoxifies drugs and poisons, breaks down hormones
Rough ER: studded with ribosomes on the outer surface of the membrane, makes secretory proteins
Nucleus: contains most of the DNA; DOUBLE MEMBRANE
Chromatin: material consisting of DNA and proteins; visible in cell division as individual condensed chromosomes
Nucleolus: nonmembranous structure involved in production of ribosomes
Nuclear envelope: DOUBLE MEMBRANE enclosing the nucleus, separating its contents from the cytoplasm, pierce by nuclear pores that serve as "holes" that regulate entry and exit of materials through envelope; continuous with ER
Bound Ribosomes: attached to the outside of the ER or nuclear envelope & make proteins for insertion into membranes
Free Ribosomes: suspended in the cytosol, proteins of these function within cytosol
Trans face: shipping department of Golgi, gives rise to vesicles that pinch off and travel to other sites
Cis face: receiving department of Golgi, located near the ER, transport vesicles used to more from ER to Golgi, ER lumen and membrane fuse with the Golgi membrane
products of the ER are modified, stored, and then sent to other destinations
Cells attach to the matrix with the help of fibronectin- binds to cell-surface receptor proteins (integrins) in the membrane
Contains collagen (glycoprotein) that forms strong fibers outside the cells
Microfilaments: thin solid rods that are built from actin molecules; help maintain cell shape, changes in cell shape, muscle contraction, cytoplasmic streaming, cell motility, and cell division
Actin-protein interactions help cytoplasmic streaming (the circular flow of cytoplasm within cell)
Actin filaments and Myosin (motor protein) interact to cause contraction of muscle cells
Intermediate filaments: maintain cell shape, anchorage of organelles, and help with formation of nuclear lamina
Microtubules: hollow rods constructed from globular proteins called tubulins; maintain cell shape, cell motility in cilia and flagella, chromosome movements in cell division, and organelle movements
Dyneins have two "feet" that "walk" along the micotubules and use ATP to cause bending movement
Large motor proteins called dyneins are attached along each outer microtubule double and are responsible for the bending movement of flagella and cilia
Inner membrane: coiled with infoldings (cristae) and divides the mitochondrion into TWO internal compartments
Mitochondrial Matrix: enclosed by the inner membrane, contains different enzymes and mitochondrial DNA and ribosomes
Intermembrane Space: narrow region between the inner and outer membrane
Outer membrane is SMOOTH
Autophagy: lysosomes fuse with vesicle containing damaged organelles and hydrolytic enzymes are used to recycle the cell's own organic material
Phagocytosis: lysosomes fuse with a food vacuole and hydrolytic enzymes digest the food particles
excessive leakage from A LOT of lysosomes leads to self-digestion of cell
Sites where DNA Replication Occurs
Each cell has an ORI (origin of replication) where DNA replication begins; prokaryotes have one ORI and eukaryotes have many
1) Helicase: enzyme that untwists the 2 strands at the replication fork
2) Single-stranded Binding Proteins: bind to the unpaired DNA strands and keep them from re-pairing
3) Primase: enzyme that synthesizes a complementary RNA primer using the parent strand as the template; new DNA strand will start at the 3' end of the primer
4) DNA Polymerase III: enzyme that adds a DNA nucleotide to the 3' end of the primer and continues to add complementary DNA nucleotides to the template strand
5) DNA Polymerase I: removes RNA nucleotides of primer from 5' end and replaces them with DNA nucleotides
6) DNA Ligase: joins final nucleotide of DNA replacement to the first DNA nucleotide of the adjacent Okazaki fragment
Nuclease: a DNA-cutting enzyme that cuts a segment of DNA strand that contains damage; the gaps are filled with nucleotides using undamaged strand
Sliding clamp: converts DNA Poly. III from being distributive (falling off) to processive (staying on)
Replicates the leading strand 5' to 3' direction; continously; only one primer Replicates the lagging strand away from replication fork, 5' to 3' direction, and discontinously synthesized in a series of Okazaki fragments, who each need a primer
Adds each monomer through dehydration reactions; is able to proofread its own work
Needs a primer & can only replicate 5' to 3' end
Topoisomerase: enzyme that helps relieve the strain by breaking, swiveling, and rejoining DNA strands
Subtopic
Stroma: fluid outside the thylakoid, contains the chloroplast DNA, ribosomes, and enzymes
Membranes of the chloroplast divide the chloroplast into 3 compartments: intermembrane space, stroma, and thylakoid space
Thylakoids: another membranous system inside chloroplasts in the form of flattened, interconnected sacs; thylakoids are stacked; each stack is called a granum
Secondary Cell Wall: between membrane and primary cell wall; has strong, durable matrix for cell protection and support
Primary Cell Wall: thin and flexible wall