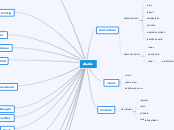
Homology
Domain/motif
Pairwise sequence alignment
MUTATIONS
Mutability
low mutability
Tyr, Phe, Leu, Cys, Trp
high mutability
Asn, Ser, Asp, Glu
can happens by
deletion
insertion
substitution
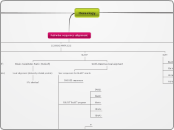
SCORING MATRICES
BLAT
more than 95% similarity
design to rapidly align longer nucleotide sequence
500 times faster than BLAST
BLAST-like alignment tool
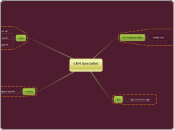
BLAST
Smith-Waterman local alignment
four component for BLAST search
CHOOSE optional parameters
output format
word size
expect value
E = Kmn e-λS
Extreme Value Distribution (EVD)
substitution matrix
turn filtering on/off
organism
different for every blast program
CHOOSE database
pnv_nr
Subtopic
pat
Pdb
RefSeq
Swissprot
Month
nr
SELECT BLAST program
tblastx
tblastn
blastx
blastn
blastp
CHOOSE sequences
Blocks Substitution Matrix (Henikoff)
local alignment (distantly related protein)
X% identical
PAM
Percent Accepted Mutations (Dayhoff)
global alignment (closely related protein)
X% divergence
PAM1
probability of occurance
+
percent accepted mutations
=
mutation probability matrix
one PAM (unit of evolution) =
1% amino acids changed between two proteins
at least 85% identical
SCORES
scores
bit scores (normalized scores)
raw scores (calculate from substitution matrix)
gap penalty
extension - elongate gap
opening - inroduce gap
mismatch
identity
Functions