ATP is present in a cell
Cellular respiration
Breaking down of glucose, and converting chemical energy from nutrients and oxygen molecules and turning it into ATP.
4 stages
Krebs cycle/ Citric acid cycle
Location: Mitochondrial matrix
Input:
2 Acetyl CoA
Output:
6 NADH, 2 FADH2, 2 ATP
Net products
6 NADH, 2 FADH2, 2 ATP
Pyruvate Oxidation
Location: Cytosol, then mitochondrial matrix
Input:
2 pyruvate, 2 CoA
Output:
2 Acetyl CoA, 2 NADH
Net products:
2 Acetyl CoA, 2 NADH
Doesn't produce ATP
Oxidative Phosphorylation
Location: Proteins in inner membrane
Input:
Oxygen gas, 10 NADH, 2 FADH2
Output:
Water, 30-32 ATP
Net products:
Water, 30-32 ATP
Produces ATP through the coupling of Chemiosmosis with electron transport chain
Electron transport chain:
Electron transport and pumping of protons (H+),which create an H+ gradient across the membrane.
Chemiosmosis:
ATP synthesis powered by the flow of H+ back across the membrane
Hydrogen ions are pumped to
intermembrane space
Glycolysis
Location: Cytosol
Input:
1 Glucose, 2 ATP
Output:
2 pyruvate, 2 NADH, 4 ATP
Net products:
2 Pyruvate, 2 NADH, 2 ATP
Produces ATP with substrate level
phosphorylation
There are 6 tyrosine amino acids and they each need a phosphate group to activate so the kinase removes...
Substrate level phosphorylation:
Production of ATP or GTP through the direct addition of a phosphate group to ADP or GDP
signal molecule - a hydrophilic molecule released by a cell to jump start a tyrosine kinase receptor
signal molecules goes to binding site
Signal binding site ( receptor) - the place where a signal molecule binds to the kinase receptors in the membrane
Inactive tyrosine kinase region - the tyrosine portion of the polypeptide that hasn't been phosphorylated.
Signal molecule binds to the kinase receptors
Dimer - when the two polypeptides that have a total of 6 tyrosine portions come together
Active tyrosine kinase region - its activated just because a dimer is formed. However it is not fully activated unless phosphate groups are attatched to the tyrosines.
Kinase - an enzyme that removes one phosphate group from ATP
ATP - Adenosine tri-phosphate. The main form of how a cell carries energy
kinase removes a phosphate group from ATP and forms...
ADP - Adenosine di-phosphate. ATP with a phosphate group removed
6 phosphate groups - the result of the kinases removing a phosphate group from each ATP
Tyrosine - an amino acid that needs a phosphate group attachment to activate
Fully activated tyrosine kinase region - when the tyrosine kinase is now able to induce a response from the cell
Now that the tyrosine kinase is fully activated inactive proteins come to it
Inactive relay proteins - proteins that relay a messages but aren't doing their job because they haven't been activated.
After interacting with the tyrosine kinases the inactive relay proteins receive a message so they become...
Active relay proteins - the relay proteins are now ready to relay a message after they've been activated.
After a long process of interacting with different relay proteins the cell can now induce a...
Cellular response - anything the signal molecule wants to the cell to do after the long events of protein interactions.
As soon as it starts, cAMP is converted to AMP
cAMP
Formed from ATP using Adenylyl Cyclase
cAMP binds to protein Kinase
Undergoes Domino Effect with other Kinase proteins
The last protein that activates brings the response form the cell.
and changes shape
G protein
GDP is replaced with GTP
G protein is now activated with GTP
binds to an enzyme
changes shape and activates
cellular response
Floating topic
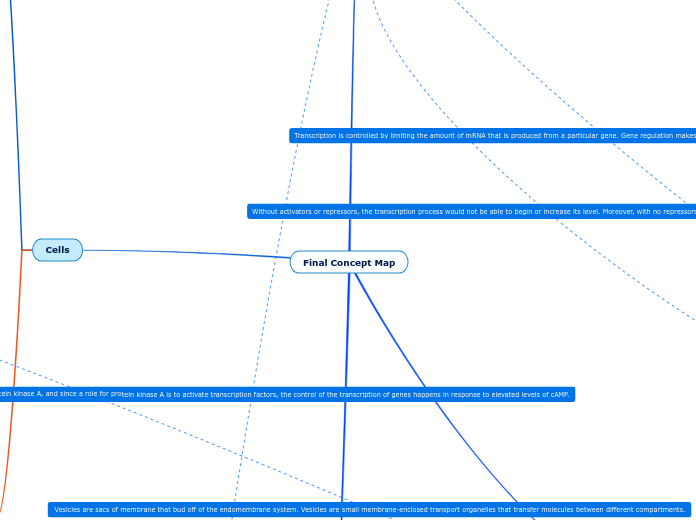
Eukaryotic Cell Gene Regulation
Transcription Stage
bound activators come in contact with mediator proteins
mediator proteins interact with proteins at the promoter
protein-protein interaction help position the initiation complex on the promoter
the repressor/activator interacts with the RNA poly II to help reduce/increase transcription rates.
activator/repressor elements come the last activated molecule activated in transduction
Pre mRNA
control elements
critical for precise regulation of gene expression
distal control elements
enhancers
sequences upstream and downstream of gene
maybe close or far from the gene they control
bin specific transcription factors (activators/repressors)
proximal control elements
sequences closer to the promoter
bind general transcription factors
Transcription factors
Specific
like activators and repressors, they bind to distal control elements called enhancers
bring about increase levels of transcriptions (activators)
bring low levels of transcription (repressors)
enhancer sequences can be present close to the gene they control or on the other side.
General
bind to the promoters and regions near the promoter to bring about basal or background level of transcription
Activators / Repressors
Activation
Repressor
Repressor proteins binds to the distal region of a transcription factor
This will block the attachment of RNA polymerase to the promotor.
This shuts down transcription
Activators
Activator proteins binds to the distal region of a transcription factor
DNA bending proteins come and bring the activators closer to the promoter
The activators bind to mediator proteins and general transcription factors
an active transcription initiation complex is formed on the promoter
How they're created
A signal molecule comes
into the cell
Binds to the receptor
on the cell membrane
A chain reaction of
secondary messengers
occurs
The last signal molecule serves as a activator or a repressor in a transcription factor
DNA binding
limits transcription
increases transcription
high level
gene expression
without repressor
operon ON
with repressor
without activator
operon OFF
with activator
operator-
the "switch
Amylase
Hydrolyze glyosidic bonds (digestive)
Albumin
Transport hormones and maintain osmotic pressure in blood
Collagen
Fibrous protein that is a support structure for cells and makes up bones, ligaments, tendons, and cartilage
Insulin
Peptide hormone that regulates blood sugar levels
Protein Synthesis
Function
Examples
Used to maintain the various functions of the cell. Proteins have a variety of functions such as being enzymes for different reactions, acting as transport proteins in the cell membrane, acting as signaling molecules, acting as antibodies and more.
Destination
Protein destination depends on
the sequence of amino acids
Protein for outside the cell
The Ribosome making the protein is bound to a signal recognition particle
The ribosome SRP complex is now bound to the rough endoplasmic reticulum.
The SRP leaves and the ribosome continues with protein synthesis.
The signal enzyme is cleaved by an enzyme in the rough ER which causes the ribosome to break apart leaving the protein alone.
The protein is then folded into its final shape
The protein is then moved into the Golgi apparatus
From the Golgi apparatus the protein is packaged and shipped to wherever it's needed in the body
Protein for inside the cell
The proteins are made useful in Eukaryotic cells with organelles such as mitochondria, Chloroplast, peroxisomes, and the nucleus
Pathway
After transcription the pre
mrna is formed
The mrna is then formed
The trna with the correct anti codon matches with the codons in mrna in the ribosome
The ribosome uses its EPA site to guide the trna and its proteins until a release factor stops the process
The stacked up amino acids from trna forms a protein
Methionyl tRNA
Formyl methionyl tRNA
3 factors
12 factors
Four factors
Three release factors
One release factor
Longer half-life(few hours to a few days)
Short half-life(seconds to few minutes
A- tRNA
Each codon gets translated and corresponding amino acids are added, all linked by peptide binds
Termination
Stop codon
Ribosome
Polypeptide chain separates from tRNA
Chain gets released
Translation
Turning of mRNA to amino acid
Asynchronous with translation
Synchronous with transcription
mRNA
Initiation
Recognition of the start codon (5'AUG)
Binding of ribosome subunits
Elongation
P- site & A- site
P- peptide chain
Transcription
Eukaryotes
Nucleus
PreRNA & mRNA
Prokaryotes
mRna
DNA template
+1 Transcription start point
INITIATION
addition of transcription factors (proteins)
RNA polymerase
RNA polymerase II
RNA polymerizes bind to the promoter and make a new strand of mRNA in the 5' to 3' direction.
ELONGATION
the RNA polymerase moves downstream adding RNA nucleotides elongating the RNA transcript in the 5' to 3' direction.
TERMINATION
RNA transcript is released and the polymerase is released form the DNA
5'cap & 3'polyA sequences
Exons & Introns
introns help to alternate spacing
RNA Processing
introns are removed to bind exons
introns are released by RNA complex and Spliceosome protein.
hydrogen bonds-
connect nitogenous bases
only in RNA
only in DNA
Uracil
Cytosine
Thymine
Guanine
Adenine
Cytoplasm
RNA
phosphodiester bonds-
connects nucleotides
pentose sugar
nitrogenous base
nucleotides
DNA
EXTRACELLULAR COMPONENTS
Cell Wall
Plasmodesmata
ECM (Extracellular Matrix)
Tight junctions
Desmosomes
Gap junctions
Cytoskeleton
Membrane Proteins
Cholesterol
Phospholipids
Fibers
Microtubules
Organelle movement
Protein Kinesin
Vesicle
Grow in centromes and pair in centrioles
Help chromosome movement
Cilia & Flagella
Intermediate Filaments
Nuclear lamina
Keratin Family
Microfillaments
Plant Cells
Central Vacuole
Use to move nutrients
Provide Support
Muscle contraction
Myosin & Actin Protein interaction
ATP
Amoebiod movement
actively (needs energy)
bulk transport
pinocytosis
endocytosis
phagocytosis
electrogenic pumps
low to high concentration
(against concentration gradient)
passively (no energy)
osmosis
diffusion of free water
facilitated diffusion
diffusion aided by
protein channels
(simple) diffusion
membrane fluidity
Lysosomes - Hydrolyzes
macromolecules
Chloroplast - Made up of
thylakoids which converts sunlight
energy to chemical energy
Centrosomes - Where cell microtubules are initiated. Contains centrioles.
Flagellum - Made up of microtubules and help the cell move (only in some animal cells)
Microvilli - projections that gives the cell more space
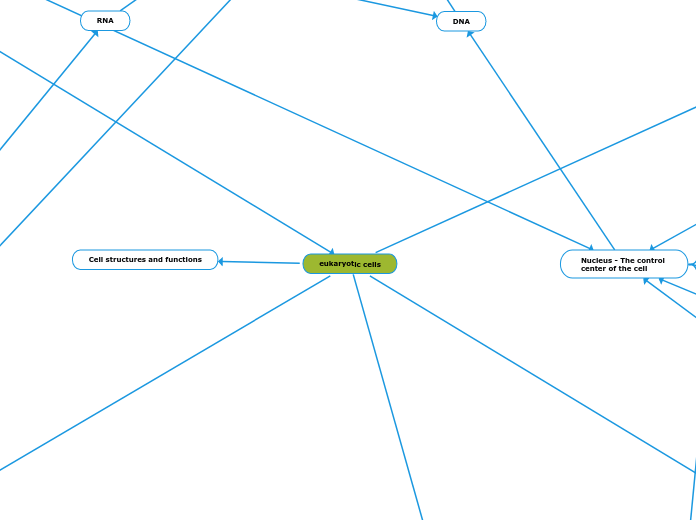
eukaryotic cells
Prokaryotic cells
Gene Regulation
operons-
cluster of functionally
related genes
two types of regulation
negative regulation
repressor-
regulatory protein
methionine(MetJ)
positive regulation
activator-
regulatory protein
CAP protein
Lack membrane bound structures
2 domains
Archaea
Bacteria
Structures
Cell wall:
Maintains cell shape, protects the interior of the cell, and prevents bursting of the cell after taking up water.
Contains peptioglycan
Plasma membrane
Surrounds the cytoplasm , separating it from the environment outside.
Ribosomes
Protein sythesis
Nucleoid:
Holds cells DNA.
Includes Proteins and enzymes that transcribe DNA & RNA that also aid in cell growth .
Cell structures and functions
Eukaryotic Animal cell structure
Eukaryotic Plant cell structure
Central Vacuole - A large organelle in plant cells. Functions as a storage, hydrolysis of big molecules, and breaks down waste products
Cell wall - Made up of proteins,
cellulose, and polysaccharides.
It protects the cell from
damage and maintains the
cell shape.
Plasmodesmata - Connects the
cytoplasm of one cell to an
adjacent cell by breaching
the cell wall
Cell membrane - Made up of a phospholipid bilayer and controls what enters and exits the cell.
proteins
carbohydrates
extracellular matrix (ECM)
phospholipids
glycerol
hydrophilic fatty acid head
hydrophobic fatty acid tails
saturated
fluid and not tightly
packed (good)
unsaturated
viscious and tightly
packed (bad)
cholesterol
regulates movement
of phospholipids
is selectively permeable
Membrane receptors
G protein coupled receptor
signal molecule
G protein coupled receptor (GPCR)
GPCR becomes activated
Tyrosine Kinase Receptor
molecule comes from another cell
Ribosomes - They bind to mRNA to synthesize proteins
Cytoskeleton - Made up of microfilaments, Intermediate filaments, and microtubules to reinforce the cell's shape.
Peroxisomes - a membrane bound organelle that produces hydrogen peroxide and then converts it to water
Golgi Apparatus - packages proteins and lipids to be transported out of the cell
Mitochondria - Makes ATP for the
cells and cellular respiration
occurs here
Endoplasmic reticulum - Made up of membrane sacs and tubes which synthesize, folds, and transport proteins
Smooth ER - Does not have
ribosomes so it's appearance
is smooth and it synthesizes
lipids and phospholipids.
Rough ER - Filled with bound
ribosomes that produces
proteins for the cell
Nucleus - The control
center of the cell
Chromatin - Mixed DNA and protein material
Nucleolus - Produces ribosomes in the nucleus
Nuclear Envelope - Membrane
filled with pores around
the nucleus. Controls
what enters and exits the nucleus.